Biographical information

I am lecturer in the School of Mathematical Sciences, University of Nottingham (my University web page is here). Before this, I was a an RCUK Academic Fellow and a EPSRC Statistics Mobility Fellow, working with Andy Wood on new methodologies for statistical problems involving shape and diffusion data.
My interests remain focused on statistical problems involving data that lie on manifolds (shape data being one example) and on inference for dynamical models.
Previously, I studied at Nottingham for undergrad (MMath, 1997-2001) and postgrad (PhD, 2003-2006) degrees in Mathematics, the latter under the supervision of Sarah Waters and Oliver Jensen.
Research interests
Manifold data
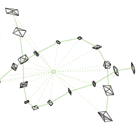
Examples of manifold data include directional data, axial data, and shape data.
Shape can be studied by identifying the position coordinates of key features of an object (landmarks) then identifying shape with all the information contained within these coordinates that is invariant under translation, rotation and scaling.
One approach to shape analysis is to work with a unit-scaled quadratic form, related to multi-dimensional scaling, of the landmark data, a form that satisfies the invariance properties, leads to intuitive definitions of mean shape, and enables computations which are fast enough to make feasible the development of bootstrap inference procedures.
Some MATLAB code for performing classical shape analysis, together with a tutorial on how to use it, is on the Shape Analysis page.
Kume, A., Preston, S.P. and Wood, A.T.A. - Saddlepoint approximations for the normalizing constant of Fisher-Bingham distributions on products of spheres and Stiefel manifolds, Biometrika, 2013.
Preston, S.P. and Wood, A.T.A. - Bootstrap inference for mean reflection shape and size-and-shape with three-dimensional landmark data, Biometrika, 2011.
Preston, S.P. and Wood, A.T.A. - Two-sample bootstrap hypothesis tests for three-dimensional labelled landmark data, Scand. J. Stat., 2010.
Inference for dynamical models
Differential equations are often a natural choice for modelling phenomena in biology, ecology, physics and many other fields. Usually they are simple to solve, but the inverse problem - of making inference about models on the basis of experimental data - is much harder. The challenge usually comes from a strongly non-linear relationship between the observed variables and the model parameters.
Stochastic differential equations and Markov random jump processes present an even tougher inferential challenge: only in very simple cases is the likelihood available analytically, meaning simulation and approximation methods are needed.
White, S.R., Kypraios, T., Preston, S.P. - Piecewise Approximate Bayesian Computation: fast inference for discretely observed Markov models using a factorised posterior distribution, 2013 (to appear in Statist. Comput.)
Preston, S.P. and Wood, A.T.A. - Approximation of transition densities of stochastic differential equations by saddlepoint methods applied to small-time Ito-Taylor sample-path expansions, Statist. Comput., 2012
Bioinformatics
Techniques of multivariate analysis are well suited to many types of data arising from new high-throughput methods in the biosciences. I am interested in how techniques such as multidimensional scaling can be further developed to deal with issues such the vast size of the datasets, having data of different types from multiple experiments, and having missing observations.
Cell motility and mechanics
Motility of immune cells is fundamental to the immune response: an important example is how T cells performing apparently random walks within lymph nodes in search of antigen-presenting dendritic cells. If immune surveillance indeed depends on this stochastic mechanism then it leads to some interesting quantitative questions.
Some cells are able to sense when they are under physical stress and change their behaviour in response - a phenomenon called mechanotransduction. Trying to understand how a cell bears a load motivates looking at the mechanics of simple cell analogues (vesicles) under various modes of deformation.
Preston, S.P., Waters, S.L., Jensen, O.E., Heaton, P.R., and Pritchard, D.I. - T-cell motility in the early stages of the immune response modelled as a random walk amongst targets. Phys. Rev. E, 2006. link
Preston, S.P., Jensen, O.E. and Richardson, G. - Buckling of an axisymmetric vesicle under compression: the effects of resistance to shear. Quart. Jl Mech. Appl. Math., 2008. link